Time Traveler
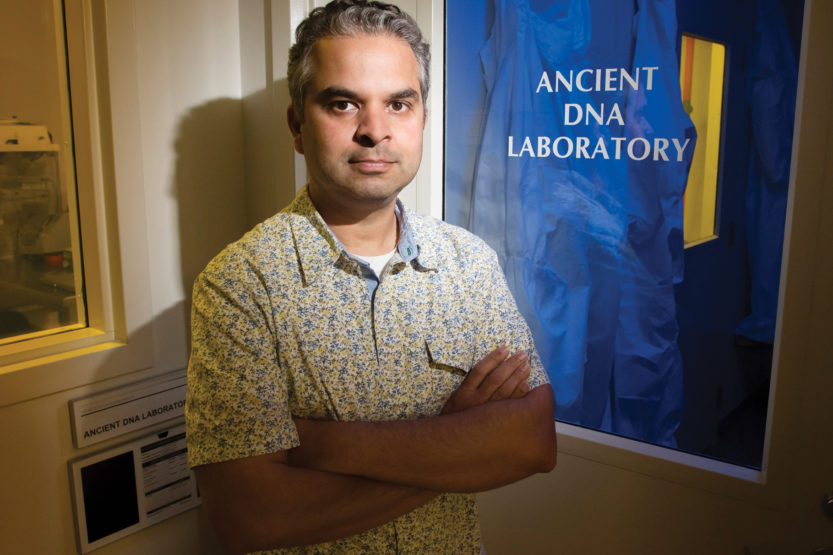 Ripan Malhi, associate professor, UI Dept. of Anthropology, focuses his research on Native American populations, particularly of the Pacific Northwest. He strongly believes in establishing relationships with tribal communities, and shares his findings and updates with them. (Image by L. Brian Stauffer)
Ripan Malhi, associate professor, UI Dept. of Anthropology, focuses his research on Native American populations, particularly of the Pacific Northwest. He strongly believes in establishing relationships with tribal communities, and shares his findings and updates with them. (Image by L. Brian Stauffer) Who says dead men tell no tales? Consider Shuká Káa, a young man who met his demise more than 10,000 years ago while hunting a bear. Thanks to anthropologists and geneticists, Shuká Káa has been able to tell us a great deal. Enormous advances in genomic science have made possible the seemingly impossible—unlocking the genetic code in ancient bones. At the University of Illinois, Ripan Malhi holds keys to the human history locked up in DNA—those molecules, present in the cells of all living organisms, that deliver unique instructions for growth, development and reproduction. A pioneer in the evolving field of molecular anthropology, Malhi is on an extended journey back in time, in quest of information about long-ago humans and the lives they led.
“Genomic analysis is now extremely informative,” says Malhi, whose easy smile and friendly manner belie his intense attention to detail. “Especially these days, when you can analyze complete genomes, not only in living people, but in ancient people as well.”
Such analyses, Malhi says, yield new data about the lifeways of peoples of the past and how they moved around the world and adapted to different environments. “Then,” he says, “we can try to combine genomic information with archaeological information and (at least in the Americas) indigenous oral histories to further enrich our understanding of the past.”
Malhi focuses much of his research on Native American populations, particularly those of the Pacific Northwest. His groundbreaking comparisons of human DNA from the present and from long ago suggest that tribal communities there include living descendants of ancient individuals. In the process, he has built strong and respectful ties to those communities.
His most recent study includes an analysis of the bones of Shuká Káa (in Tlingit, “Man Ahead of Us”), which were discovered in 1996 in the On Your Knees Cave on Alaska’s Prince of Wales Island and date back more than 10,000 years. DNA from three other individuals found along the coast of British Columbia within 200 miles of the Alaska site also are part of the study; those remains date back 1,750, 2,500 and 6,075 years, respectively. (Control samples from ancient individuals found in Washington state and Montana also were included.)
The study shows genetic continuity between those ancient individuals and 100 members of six communities in the Pacific Northwest, including Metlakatla and Lax Kw’alaams communities on Prince of Wales Island and Tlingit- and Haida-speaking people in southeast Alaska. The work, which also involved researchers from the University of Chicago, Penn State, the Sealalska Heritage Institute and the University of Oklahoma, was published in April 2017 in the Proceedings of the National Academy of Sciences.
“Indigenous communities on the Northwest Coast have oral histories that talk about how they have been in that region since—they call it ‘time immemorial,’” Malhi says, noting that his study supports that tradition, establishing “genetic continuity from the present day back to 10,000 years ago.”
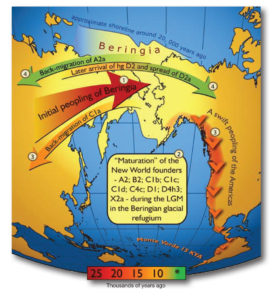
Native American oral histories underpin the theory that humans first entered North America from Asia, crossing a glacial land bridge called Beringia. Malhi’s research indicates that early immigrants found progress to North America blocked by glaciers, which led them to evolve new traits. (Image courtesy of Wikiwand)
CROSSING THE BRIDGE
Those oral histories underpin the theory that humans first entered North America from Asia, crossing a glacial land bridge that spanned what is now the Bering Strait. One hypothesis, for which Malhi provided genetic evidence in a 2007 study, makes the case that early immigrants had an extensive layover en route to their new home. An enormous amount of the world’s water was locked up in glaciers at the time—the Last Glacial Maximum, which peaked in 24,500 BCE and lasted more than 6,000 years—blocking ingress to the North American continent. Malhi says his study suggests “that ancestors of Native Americans lived in isolation on a land bridge called Beringia.” Living for generations in this glacial refugium, the migrating people adapted and evolved new traits. Once the world warmed and the glaciers started to melt, the population was able to enter North America and spread rapidly across the continent. “That’s why you find mitochondrial genomes in the Americas that are unique to the Americas,” Malhi explains.
Mitochondrial DNA passes from mother to child. It is far more abundant in cells than the complete DNA in the nucleus. Mitochondrial DNA has proven extremely useful in genomic research, but it tells just half of an individual’s genetic story. Among Malhi’s achievements is his successful isolation and analysis of the nuclear DNA in ancient samples. He is among a small vanguard of scientists doing so, based on the pioneering work of a group in Denmark who mapped the first genome of an ancient individual in 2010. “Ten years ago, I would never have imagined sequencing complete genomes from ancient individuals,” Malhi says. “But now we’re doing it.”
Ancient DNA comes from a variety of sources—generally museums, universities and other institutions that house excavated remains. It is essential to obtain permission from the indigenous communities associated with the remains. The bone sample required is extremely small, and the results of the analysis may be used in multiple studies.

Malhi helped establish the Summer INternship for Indigenous Peoples in Genomics, which brings together indigenous students to study the cultural, legal, ethical and scientific aspects of genomic research. The program has drawn participants throughout the U.S., Canada and Mexico. (Image by Kathryn Flynn, Source: Ripan Malhi)
NATIVE TRUST
Strong relationships with native communities are vital to collecting living DNA samples. Community members frequently view genetic research as one more form of cultural and racial exploitation, of no benefit to them. The long, tragic history of cultural artifact removal—including human remains—to the research institutions of colonial powers substantiates such suspicions. Government regulations, both in the U.S. and Canada, now acknowledge and protect the rights of indigenous people to their heritage, including sacred sites and language, as well as human remains and archaeological artifacts.
Malhi’s field work with DNA began in 2001, when he was a doctoral candidate in molecular anthropology at the University of California, Davis. “A lot of the samples I analyzed as a graduate student were already in the lab,” he recalls. “I could have just gone to the freezer, never left the lab, never interacted with any indigenous people at all. But toward the end of my graduate career, I decided that didn’t seem quite right.” He started contacting indigenous groups in California and visited Round Valley Reservation, which is on the edge of Mendocino National Forest, north of San Francisco. People there were openly skeptical of his request to collect DNA sam-ples (typ-ically from saliva, swab-bed from the inside of an individual’s mouth).

Harold Harry (left), community member and collaborator on the Stswecem’c Xgat’tem First Nation, and Jerry Cybulski, curator of the Canadian Museum of History in Gatineau, Quebec. Malhi met Cybulski in 2004, and the two collaborated on research until Cybulski’s retirement in 2013. (Image courtesy of Ripan Malhi)
“There were community members who asked, ‘Why should we trust you?’” Malhi recalls. “They relayed stories of researchers who had collected samples, left and never came back. I realized I didn’t want to do that. But I really didn’t know the best way to proceed with research until I met Jerry.” That was in 2004. Jerry Cybulski, a curator at the Canadian Museum of History in Gatineau, Quebec, contacted Malhi who, by that time, had co-founded Trace Genetics, a startup that specialized in analyzing indigenous North American DNA. Cybulski had obtained samples from two 5,000-year-old skeletons found in China Lake in central British Columbia, and he believed the pair to be brothers—a hypothesis that Malhi’s testing proved to be true. Cybulski then arranged for Malhi to join him in giving a talk to residents of Canoe Creek, the indigenous stakeholder community where the skeletons were discovered.
“Ripan was amazing. He presented his information in a really interesting way, and it just grabbed everybody’s attention,” Cybulski says, noting that Malhi went on to keep community members involved in the project, reporting on findings and returning to give updates. “One has to be willing to visit communities and talk. And listen,” Cybulski says. “That’s what I’ve been doing my whole career. And Ripan just fits right in.” The work initiated a collaboration between the two men that lasted until Cybulski retired in 2013.
Involvement in genetic research gives indigenous community members new information about themselves and their heritage. An especially fascinating aspect of this interaction is the affirmation of time-honored creation myths. “What we’ve been finding is support for oral history,” Cybulski says. “First Nations native people have longstanding tales about their origins and where they have been at different times in their existence. ‘Since time immemorial’ may be a worn expression, but it’s true. The DNA work has been supporting that.”
EUROPEAN CONTACT
Another study led by Malhi, published in 2014, traces a significant genomic change in Northwest Native American communities. The change took place in the mid-19th century, with the arrival of European explorers and settlers. “When Europeans brought over diseases like smallpox, it altered the genomes of indigenous peoples,” says Malhi. “Variants that may have been adaptive to the previous environment before Europeans came were no longer adaptive and may actually have been detrimental. Those variants were wiped out of the population.”
Malhi and his team also modeled demographic history using the same genomic information. “There was a huge reduction in the genetic diversity of the indigenous population—close to a 60 percent bottleneck in diversity—as a result of the European contact and colonization,” he says, noting that this reduction may have been due to warfare and forced changes to a more Westernized culture as well as disease.
Malhi’s cultural sensitivity developed along with his scientific skills. At Trace, “I started really thinking about what it means to have DNA from a certain population and how that contributes to your identity,” he says. “Over the years, I’ve come to the conclusion that, yes, biology contributes to your identity. But so do history, language and culture.” He sold the company in 2005 and moved to Illinois the following year, taking a joint post in anthropology and animal biology at the University. He has since added an affiliation with the Carl R. Woese Institute for Genomic Biology, an interdisciplinary UI research center at the forefront of the field.
If Malhi is good with people, he’s superb at the bench. His molecular anthropology lab on campus, one of only a dozen such facilities in the world, is designed to prevent contamination of precious ancient samples. “Human DNA,” Malhi says, “is everywhere,” and hence the lab is, “forensic on the extreme end.” Positive air pressure repels contaminants from the outside world, and there’s room inside for just two researchers at a time, clad in sterile white coveralls known as “bunny suits.”
The bone sample required for ancient DNA analysis is astoundingly small—one-fifth of a gram, approximately the size of a tooth cavity. The sample is drilled to a powder and decalcified, separating DNA from protein. Chemical reagents break up the cells and the DNA, which is negatively charged, and then binds with a column of silica. Other materials are then washed away. The resultant gene sequences are so tiny they’re measured in trillionths of a gram. Analyses extend beyond the bone samples to dental calculus and coprolites (“poop,” Malhi explains with a grin) from the ancient individual, creating an extract that “includes not only the DNA of the person but also of all the plants and animals that person may have eaten,” he says.
The work then moves to the supercomputer. Computational and statistical analysis, using sophisticated bioinformatics software, contextualizes the DNA sequences by comparing them with complete genomes. The sequences are used to build a “genetic library” of an individual’s life and genetic heritage.
“Most of the bottleneck in doing this type of analysis used to be in the wet lab,” says Malhi. “Now, getting the data is a lot easier. The intensive part comes from analyzing the hundreds of thousands to millions of DNA sequences you get once you do the sequencing.” Malhi works with the Carver Biotechnology Center on campus “to adapt what I need to generate sequences from these ancient individuals,” he says. “Once we have the data, the Institute for Genomic Biology has a biocluster [a high-performance computing resource] where we can do intensive analyses and computational processes.”
Beyond involving stakeholder communities in the study of their genetic heritage, Malhi is recruiting members of indigenous groups to his field. The annual Summer INternship for Indigenous Peoples in Genomics, which he helped launch in 2011, brings together indigenous students to study the cultural, legal, ethical and scientific aspects of genomic research. During the week-long program, participants analyze their own DNA. Some of the students have gone far beyond this relatively brief opportunity, which is intended to recruit talented young Native Americans to STEM fields.
“The internship was an incredible experience for me,” says Nathan Nakatsuka, a Kanaka Maoli (native Hawaiian) who attended the first workshop, held on the Illinois campus in 2011. “The experience was one of the major influences that caused me to switch to population genetics research.” Now a fourth-year MD/PhD student at Harvard, Nakatsuka studies population history and medical genetics of Native Americans and the native peoples of India.
So the generations arc across the millennia, from a time when glaciers ruled the Earth to the day a hunter met his end in Alaska, to the future of a young medical student who wants to know more about what makes us human. This is the story of genomic research, and important parts of it are being written by University of Illinois Professor Ripan Malhi.
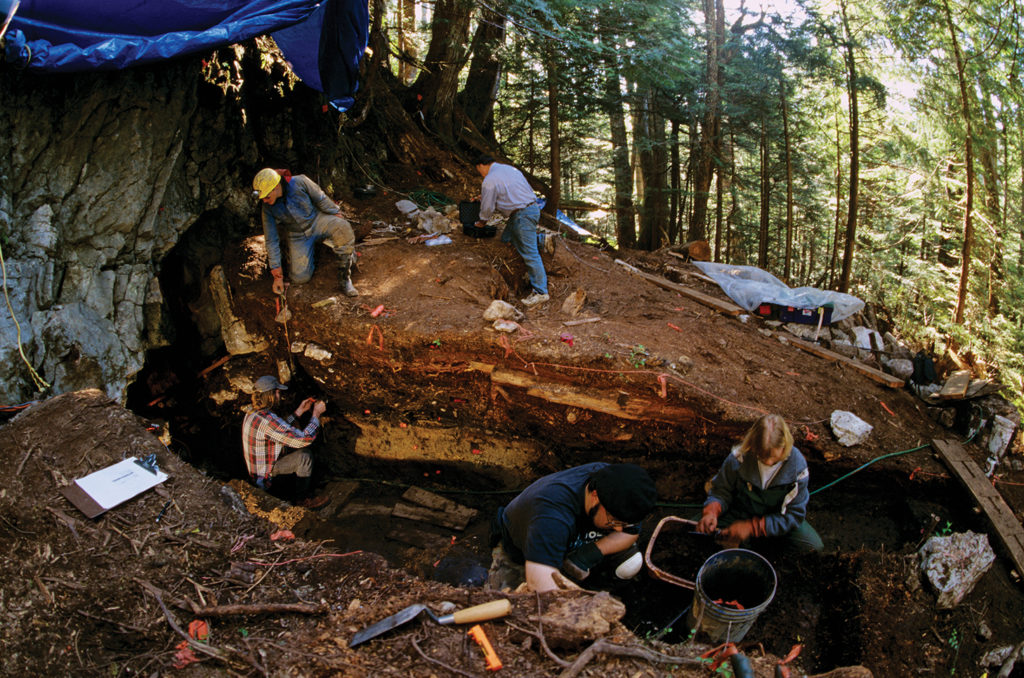

The bones of Shuká Káa were discovered in 1996 in the On Your Knees Cave on Alaska’s Prince of Wales Island. They date back more than 10,000 years. (Image by Kenneth Garrett)
The Story of Shuká Káa
Some 10,300 years after his death, Shuká Káa lives on in DNA linking him to the Tlingit people who named him “Man Ahead of Us.”
Discovered in 1996 in the On Your Knees Cave on Prince of Wales Island in Alaska, the bones of this ancient individual are the oldest human remains found in North America. Undertaken with approval of Tlingit tribal elders, who conferred Shuká Káa with his name, DNA testing revealed that he was approximately 20 years of age, had lived on seafood and was likely a mariner killed by a bear.
In 2008, his remains were ceremonially interred on Prince of Wales Island by the Tlingit. His headstone states: “We Have Lived in Southeast Alaska Since Time Immemorial. Shuká Káa Is a Testimony to Our Ancient Occupancy of This Land.” In 2017, UI genomic anthropologist Ripan Malhi (see main story) analyzed nuclear DNA from Shuká Káa and found genomic markers shared by the Tlingit and other indigenous inhabitants of the Pacific Northwest.



